There is emerging evidence that lncRNAs can be involved in various critical biological processes. However, our understanding on lncRNA is still at the rudimentary stage. Zebrafish is a full-developed model system being used in a variety of basic research and biomedical studies. Hence, it is a good idea to study the role of lncRNA using zebrafish as a model. Here, we constructed ZFLNC -- a comprehensive database of zebrafish lncRNA that is dedicated to offering a zebrafish-based platform for deeply exploring lncRNA function and mechanism, to the relevant academic community.
In order to set up a zebrafish-based platform for deep exploration of the functions and mechanisms of zebrafish lncRNAs and their mammalian counterparts, we constructed ZFLNC, which is a comprehensive database of zebrafish lncRNA with three main goals: (i) collecting the most complete dataset of zebrafish lncRNAs, with the most comprehensive annotations; (ii) Using a variety of conservation analysis methods to study the potential lncRNA orthology; (iii) providing a user-friendly website with useful web-based tools for the functional interrogation of conserved lncRNAs.
The principal data resources of lncRNAs in this database come from NCBI, Ensembl, NONCODE, zflncRNApedia and literature. We also obtained lncRNAs as a supplement by analyzing RNA-Seq datasets from SRA database. We carried out the expression profile, GO annotation, KEGG pathway annotation, conservative analysis and OMIM annotation for those zebrafish lncRNAs. In the current version ZFLNC contain 13,604 lncRNA genes and 21,128 lncRNA transcripts. To our best knowledge, ZFLNC should be the most comprehensive and well annotated database for zebrafish lncRNA.
a. Data source
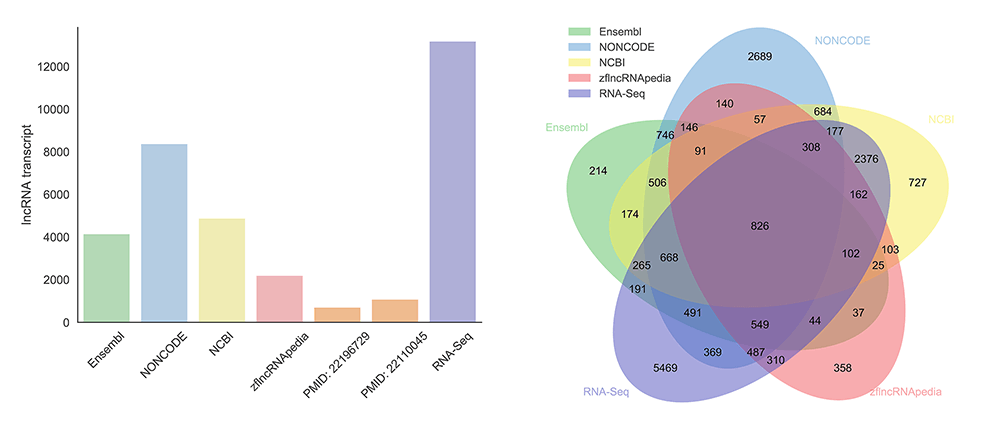
We obtained 7394 zebrafish lncRNA genes (13166 transcripts) from RNA-Seq data, and then integrated them with those from Ensembl, NONCODE, NCBI, zflncRNApedia and literature. Our final zebrafish lncRNA set contains 13604 lncRNA genes (21128 transcripts). The major data sources of our lncRNA set are from RNA-seq data analysis and NONCODE, followed by NCBI, Ensembl and zflncRNApedia orderly. Venn diagram shows that, although there is an obvious overlapping between different sources, a lot of unique lncRNAs emerged in RNA-seq datasets and NONCODE.
b. RNA-Seq Analysis
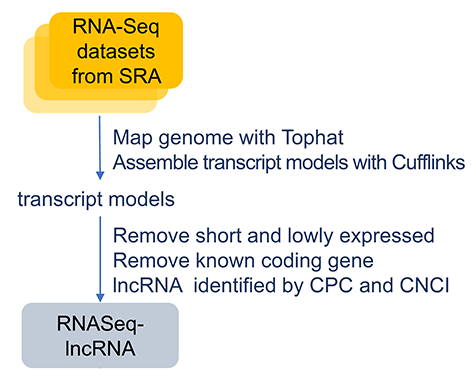
RNA-Seq data were downloaded from NCBI SRA database (RNA-Seq_from_SRA.xlsx).SRA format files were dumped to FASTQ format files by SRA-Toolkit. Low quality reads were trimmed by Trimmomatic (Version 0.32). RNA-Seq reads were mapped to zebrafish genome (Zv9) using Tophat2 (Version 2.0.13), then transcriptome was assembled by Cufflinks (Version 2.2.1). Multiple-exon transcripts were considered as being expressed if they had an FPKM greater than 0.1. For single-exon transcript, more rigorously, FPKM should be greater than 5 and transcript length greater than 2000. Those foregone coding-genes or transcripts with size less than 200nt were filtered out. Then, lncRNA candidates were identified by CPC (Version 0.9-r2) and CNCI (Version 2). At last, all zebrafish lncRNAs stemming from RNA-seq datasets and other publicly available sources were integrated together using the Cuffmerge program in the Cufflinks suite.
c. Co-expression Profiling and GO/KEGG annotation
The expression profile of zebrafish lncRNAs and coding-genes were quantified by Cuffnorm program in the Cufflinks suite and then scaled by upper-quartile normalization. We then calculated the Spearman's correlation coefficient and its corresponding P-value between the expression profiles of each gene-pair using the in-house Perl script. Only gene-pair with an adjusted P-value of 0.01 or less and with a Spearman's correlation coefficient no less than 0.5 is regarded as co-expression in our coding-lncRNA gene co-expression network.
The GO annotation of zebrafish coding-gene was downloaded from Gene Ontology Consortium (only biological process annotations were considered). While, GO annotation of zebrafish lncRNA was predicted using the goatools (version 0.6.4), which determines the GO annotation of one gene in our network according to the GO annotations of its immediate neighbor genes (P-value < 0.05).
The KEGG annotation of zebrafish coding-gene was obtained from KEGG Automatic Annotation Server using zebrafish coding-gene sequence. While, KEGG annotation of zebrafish lncRNA was predicted using the in-house Python script. The KEGG annotation of one gene in our network was determined by the enrichment of KEGG annotations according to its immediate neighborhood (p-value < 0.05), when using hypergeometric distribution.
d. Conservation Analysis
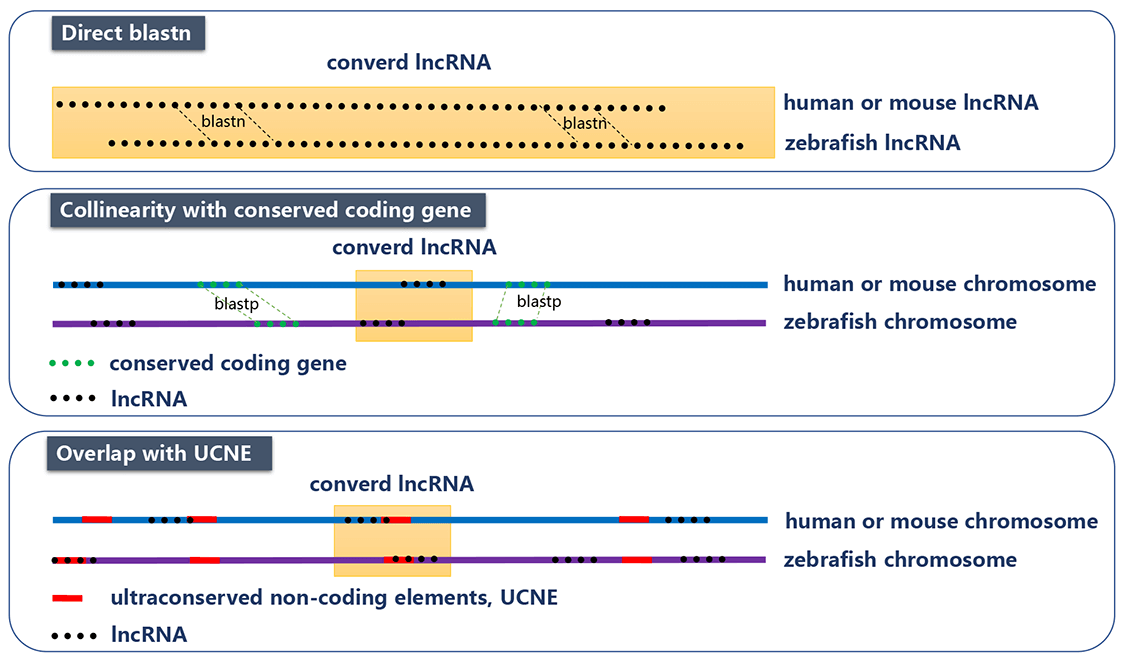
To examine the sequence conservation of lncRNAs, we used the phastCons scores calculated from the UCSC 8-way vertebrate genome alignment. We further used three methods (that are direct BLASTN, collinearity with conserved coding gene, and overlap with multi-species ultra-conserved non-coding elements (UCNE)) to find the counterparts of zebrafish lncRNAs in human or mouse. In direct comparison of zebrafish lncRNA and human/mouse lncRNA with BLASTN, bidirectional best hits using a relatively non-stringent threshold (E-value<=10-5) were considered as orthologs. In collinearity method, we compared the coding genes of zebrafish with those of human or mouse using BLASTP as anchor points. We assumed that those lncRNAs with more than 5 anchor points in the 20k upstream/downstream region are orthologs. In UCNE method, if two lncRNAs from different species overlap with at least one UCNE, as another anchor point, they were considered as orthologs. Finally, we obtained 2,156 zebrafish lncRNA genes that have the counterpart in human or mouse.
e. OMIM prediction
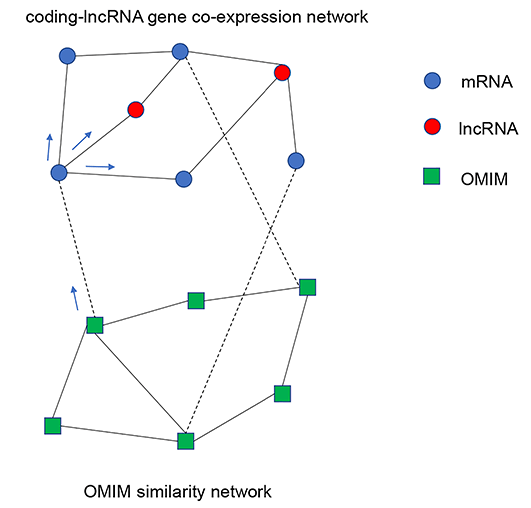
We use the RWRH (random walk with restart on heterogeneous network) algorithm to analyze the relationship between lncRNA and OMIM in MATLAB. The upper subnetwork is coding-lncRNA gene co-expression network, and the lower network is OMIM similarity network. OMIM similarity matrix is from Disimweb, and gene-OMIM relationship is from InterMine. With this approach, 291 lncRNA genes are predicted to be OMIM-related.
a. Browse
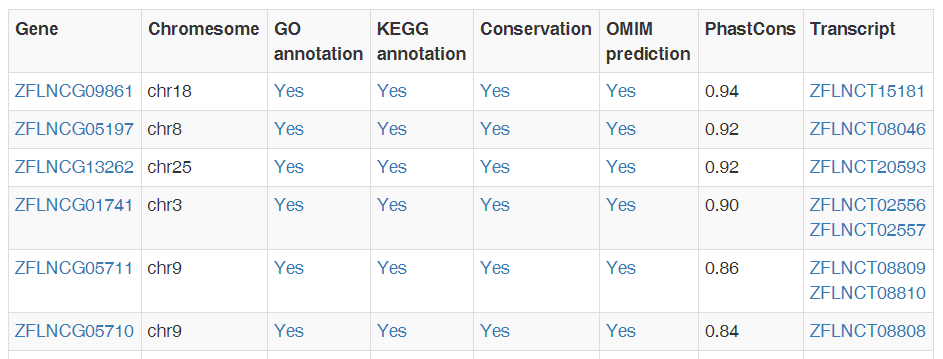
In "Browse" module, you can browse all lncRNA genes or transcripts. LncRNA is sorted according to the richness of its annotation. In particular, you can also browse lncRNAs with conservation or OMIM annotation directly in "Conservation" and "OMIM" modules.
b. GBrowser
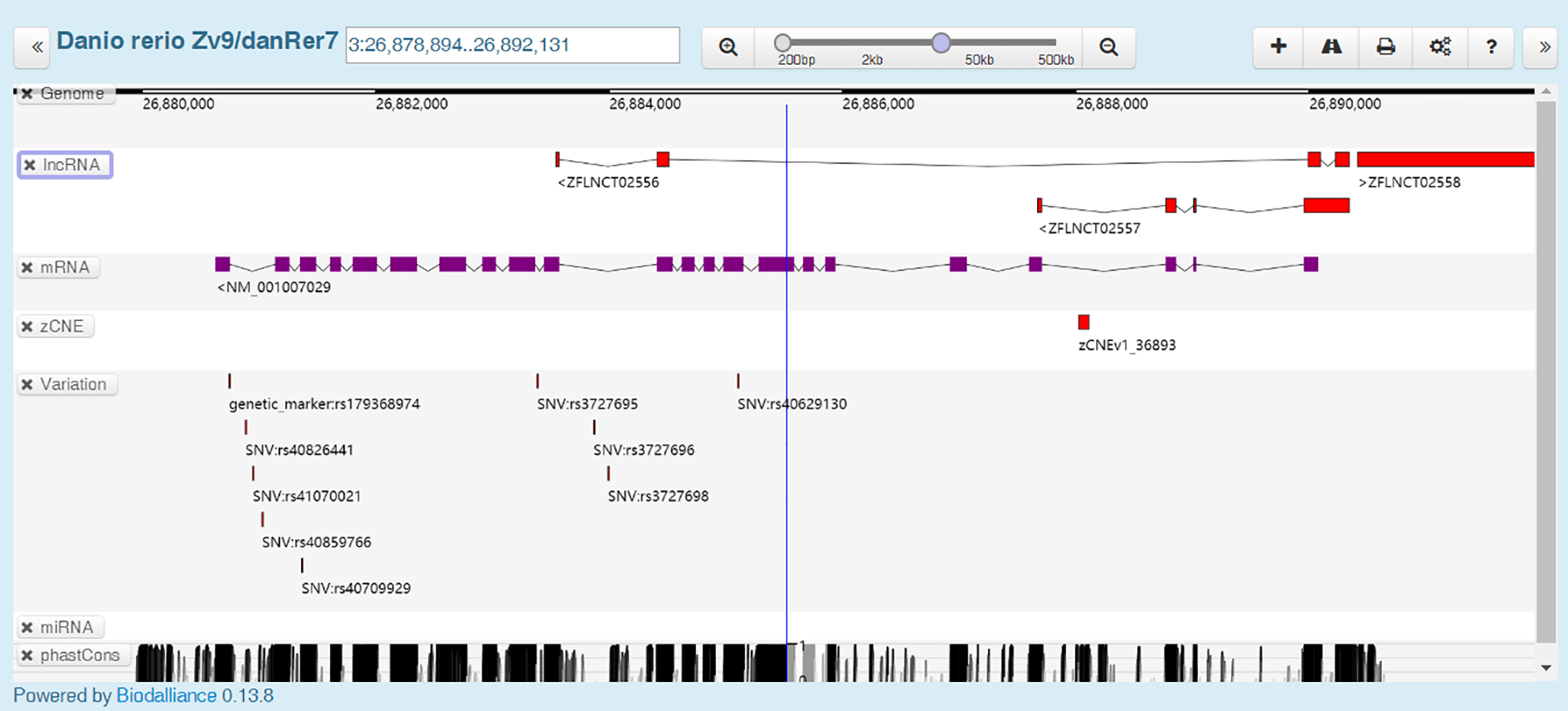
In "GBrowser" module, you can view lncRNA-related genomic annotation, such as mRNA, conserved non-genic elements, genome variation and miRNA.
c. BLAST
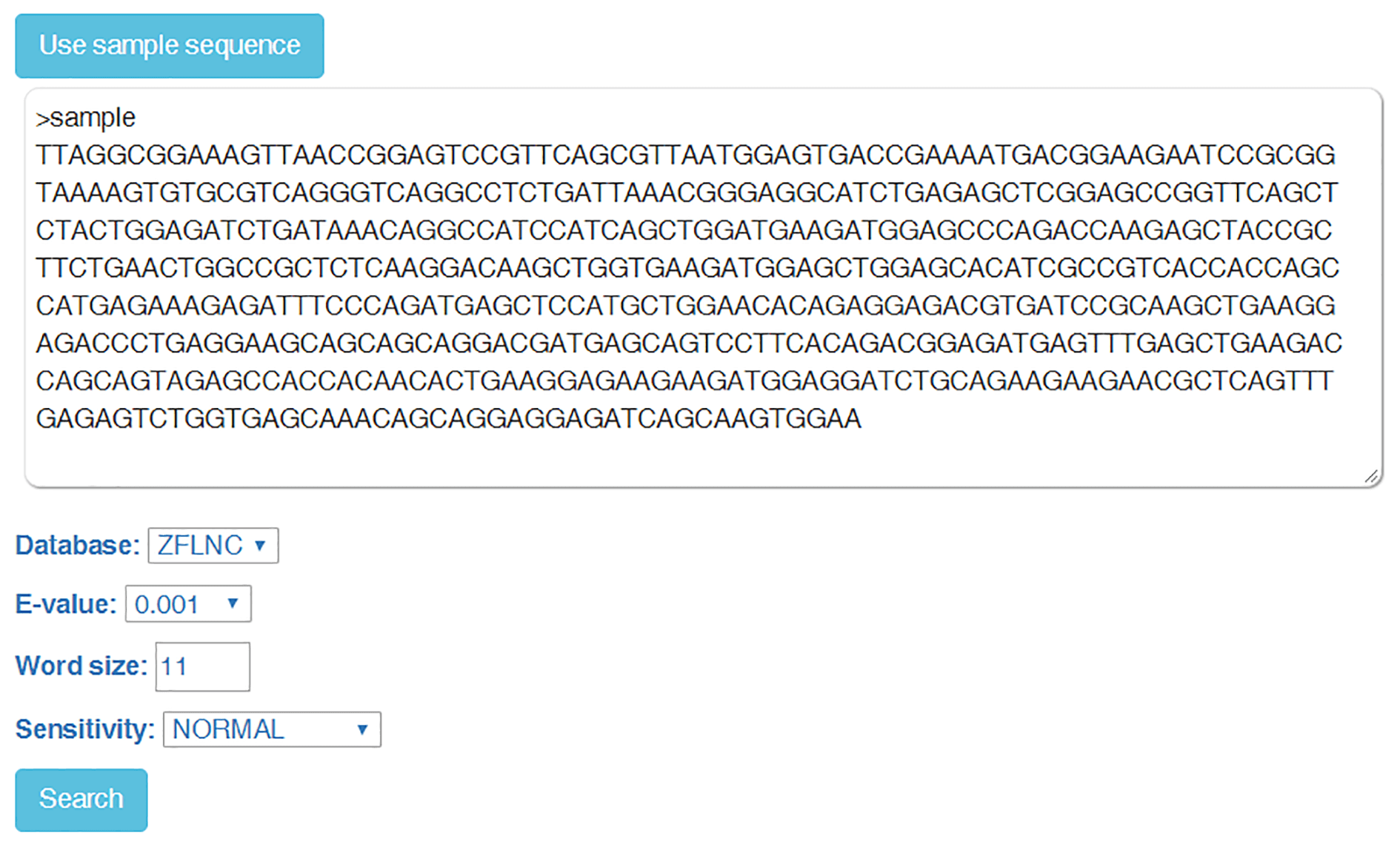
"BLAST" module can query ZFLNC based on sequence similarity.
d. ID conversion
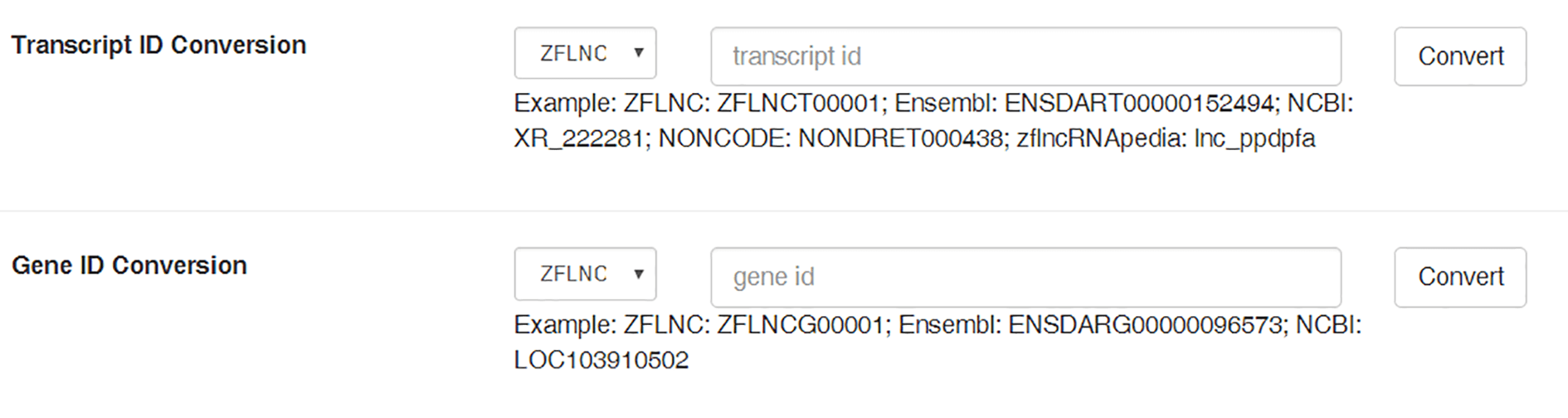
Before ZFLNC, zebrafish lncRNAs were dispersed in different databases. Hence, we provided "ID Conversion" module to facilitate the use among diverse databases. You can use BLASTN through sequence similarity, or Gbrowser through sequence positions to convert the lncRNA in other sources into ZFLNC.
5. Search
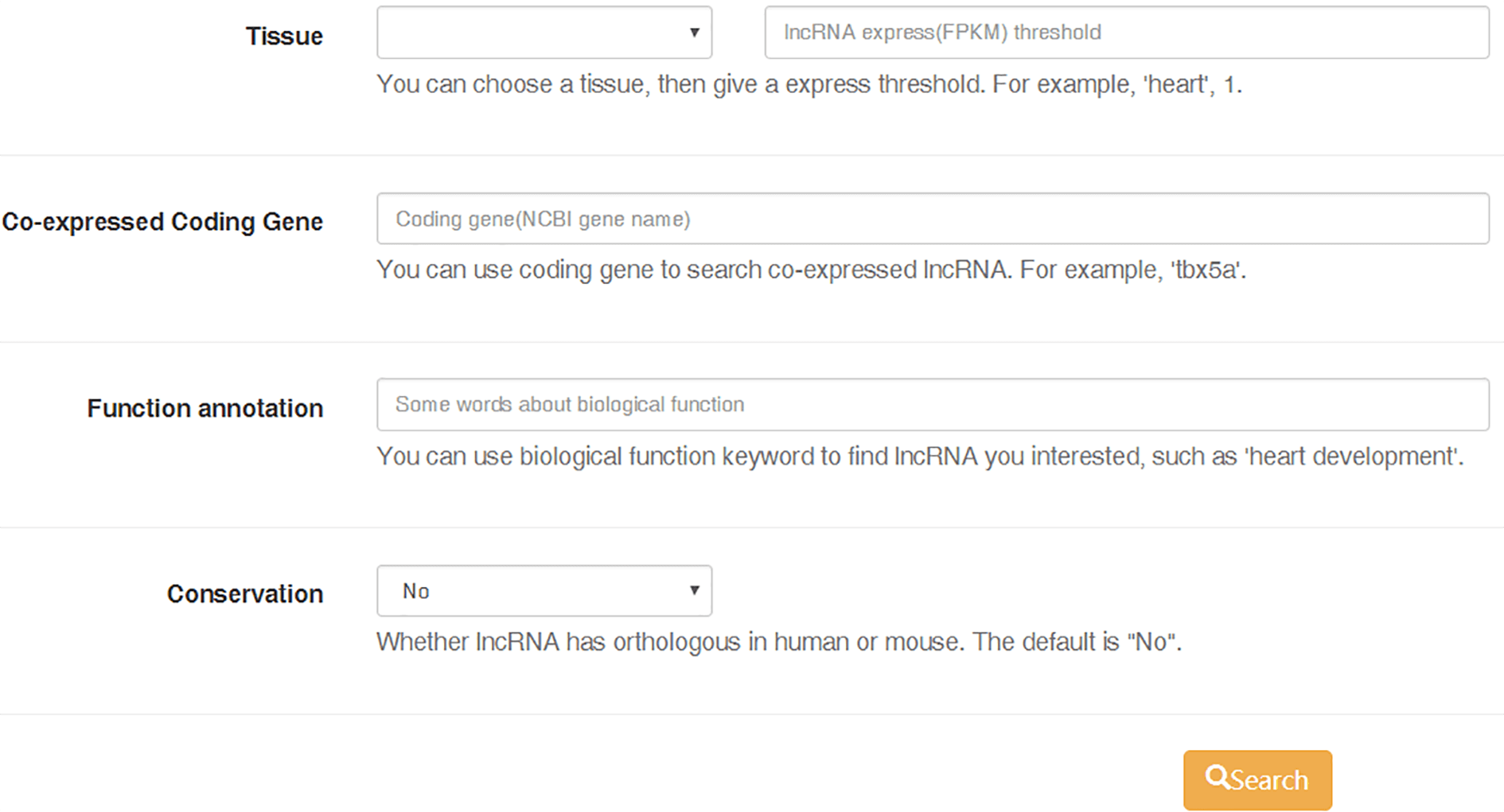
"Search" module provides a simple and fast search based on lncRNA ID, and also an "Advanced functional lncRNA filtering" function to help you find interested lncRNAs in according to its expression profile in tissue, the co-expressed coding gene and the annotated biological function (GO, or KEGG, or OMIM annotation).
